A Practical Tutorial on Knowledge Graph–Enhanced AI Retrieval (GraphRAG)
Build a production-ready tutorial for knowledge graph–enhanced AI retrieval: schema, ingestion, Cypher, hybrid search, and evaluation.
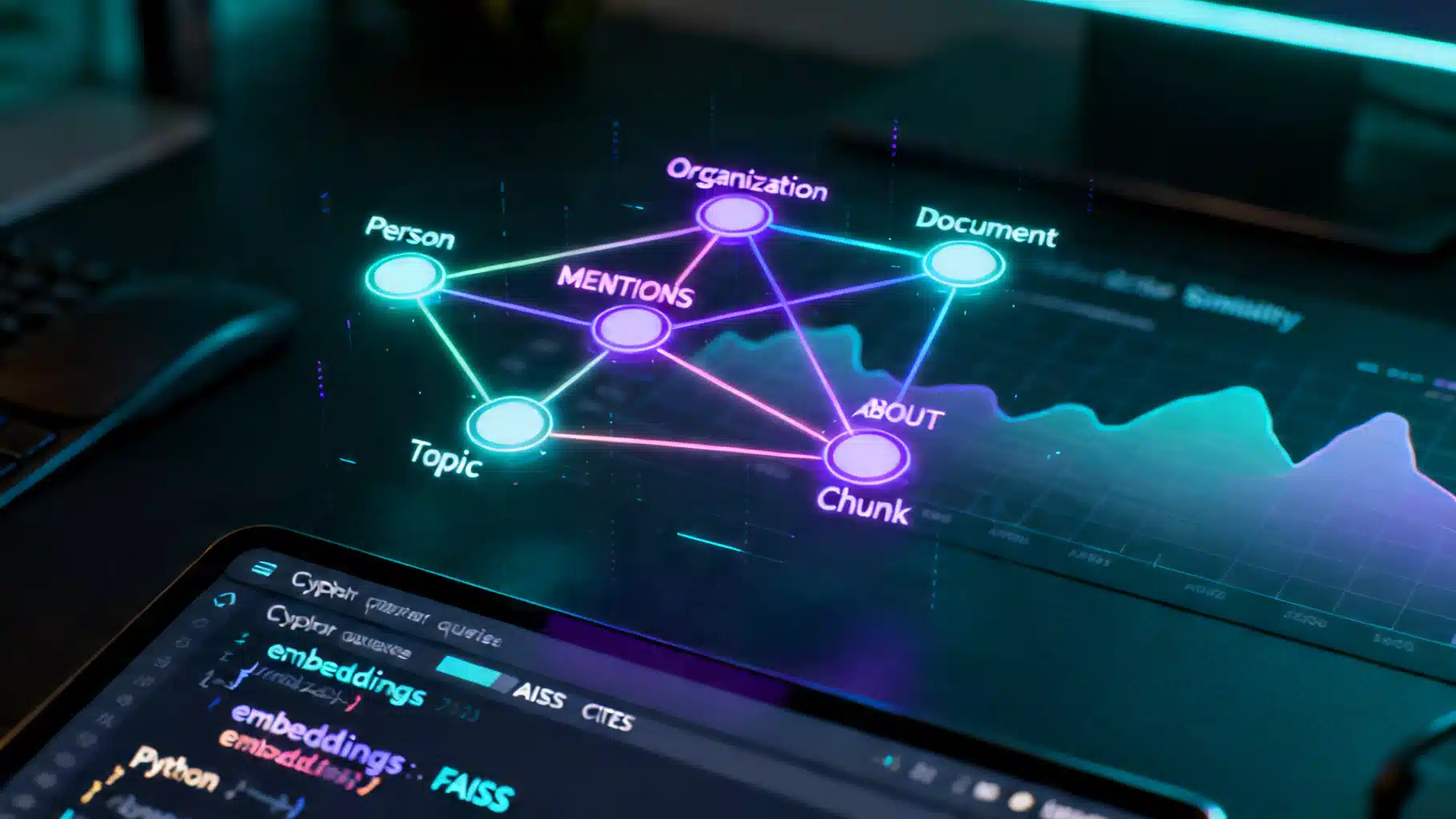

Image used for representation purposes only.
Overview
Retrieval-augmented generation (RAG) works well for fuzzy, semantic lookups. But when your data has rich structure—people, organizations, documents, events, citations—plain vector search loses context and precision. A knowledge graph (KG) restores that structure and lets you ask topology-aware questions (“Who collaborated with whom?” “What’s the provenance of this claim?”), while still benefiting from embeddings for semantic recall.
This tutorial walks you through building a graph-enhanced retrieval pipeline—often called GraphRAG—from first principles. You’ll design a schema, ingest data, create a vector index, write Cypher queries for graph traversal, and combine the results into a hybrid retriever that feeds your LLM with precise, provenance-rich context.
What you’ll build:
- A minimal KG with Documents, Chunks, Entities (Person, Organization, Topic), and typed relationships
- A vector index for semantic recall
- A hybrid retrieval function that expands from semantic seeds over graph neighborhoods and re-ranks evidence
- An evaluation harness for faithfulness and coverage
Prerequisites:
- Python 3.10+
- A Neo4j database (local or managed)
- Basic familiarity with embeddings and Cypher
Architecture at a Glance
- Ingestion: Parse documents → chunk text → extract entities/relations → upsert to Neo4j
- Indexing: Embed chunks → store vectors in FAISS (or your vector DB of choice) → map chunk_id ↔ vector_id
- Retrieval: For each question, do semantic recall (vector) → graph expansion (Cypher) → re-rank → assemble citations → generate answer
- Evaluation: Offline Q/A runs measuring faithfulness, coverage, latency, and context length
Step 1: Model a Minimal Schema
Keep the first iteration small and explicit. You can always refine.
Entities (labels):
- Person(name, aliases)
- Organization(name, aliases)
- Topic(name)
- Document(id, title, url, published_at)
- Chunk(id, text, doc_id, order, embedding)
Relationships (types):
- (Document)-[:CONTAINS]->(Chunk)
- (Chunk)-[:MENTIONS]->(Person|Organization|Topic)
- (Person)-[:AFFILIATED_WITH]->(Organization)
- (Document)-[:CITES]->(Document)
- (Chunk)-[:ABOUT]->(Topic)
Why this works:
- Chunk nodes hold the raw text you’ll feed to the LLM.
- Entity nodes capture canonical references and enable disambiguation.
- Edges preserve provenance and support topological queries (neighbors, paths, communities).
Step 2: Create Constraints and Indexes (Cypher)
Use constraints to keep your graph clean and fast.
// Neo4j 5+ style
CREATE CONSTRAINT person_id IF NOT EXISTS FOR (p:Person) REQUIRE p.name IS UNIQUE;
CREATE CONSTRAINT org_id IF NOT EXISTS FOR (o:Organization) REQUIRE o.name IS UNIQUE;
CREATE CONSTRAINT topic_id IF NOT EXISTS FOR (t:Topic) REQUIRE t.name IS UNIQUE;
CREATE CONSTRAINT doc_id IF NOT EXISTS FOR (d:Document) REQUIRE d.id IS UNIQUE;
CREATE CONSTRAINT chunk_id IF NOT EXISTS FOR (c:Chunk) REQUIRE c.id IS UNIQUE;
// Optional text indexes for name lookups
CREATE FULLTEXT INDEX entity_name IF NOT EXISTS
FOR (n:Person|Organization|Topic) ON EACH [n.name, n.aliases];
Step 3: Ingest Documents and Extract Entities
Start with a small corpus (e.g., internal docs, papers, or blog posts). Chunk by semantic boundaries (e.g., headings or ~250–400 tokens).
Python snippet using spaCy for quick NER and simple relation heuristics:
import os, uuid, json
import spacy
from sentence_transformers import SentenceTransformer
from neo4j import GraphDatabase
# Load NLP + embeddings
nlp = spacy.load("en_core_web_sm")
embedder = SentenceTransformer("all-MiniLM-L6-v2")
driver = GraphDatabase.driver(os.getenv("NEO4J_URI"), auth=(os.getenv("NEO4J_USER"), os.getenv("NEO4J_PASSWORD")))
def chunk_text(text, max_chars=1000):
parts, buf = [], []
for sent in text.split('.'):
if sum(len(s) for s in buf) + len(sent) < max_chars:
buf.append(sent)
else:
parts.append('.'.join(buf).strip()+'.')
buf = [sent]
if buf:
parts.append('.'.join(buf).strip()+'.')
return [p.strip() for p in parts if p.strip()]
def upsert(tx, query, **params):
tx.run(query, **params)
def ingest_document(doc):
# doc = {"id": str, "title": str, "text": str, "url": str|None}
chunks = chunk_text(doc["text"])
with driver.session() as sess:
sess.execute_write(upsert,
"MERGE (d:Document {id:$id}) SET d.title=$title, d.url=$url",
id=doc["id"], title=doc["title"], url=doc.get("url"))
for order, text in enumerate(chunks):
cid = str(uuid.uuid4())
emb = embedder.encode([text])[0].tolist()
ents = nlp(text).ents
sess.execute_write(upsert,
"MERGE (c:Chunk {id:$id}) SET c.text=$text, c.order=$order, c.doc_id=$doc_id, c.embedding=$emb",
id=cid, text=text, order=order, doc_id=doc["id"], emb=emb)
sess.execute_write(upsert,
"MATCH (d:Document {id:$doc_id}), (c:Chunk {id:$cid}) MERGE (d)-[:CONTAINS]->(c)",
doc_id=doc["id"], cid=cid)
# Upsert entities + MENTIONS
for e in ents:
if e.label_ in ["PERSON", "ORG"]:
label = "Person" if e.label_ == "PERSON" else "Organization"
sess.execute_write(upsert,
f"MERGE (n:{label} {{name:$name}})", name=e.text)
sess.execute_write(upsert,
"MATCH (c:Chunk {id:$cid}), (n {name:$name}) MERGE (c)-[:MENTIONS]->(n)",
cid=cid, name=e.text)
Notes:
- For production, replace simple heuristics with robust IE (e.g., pattern libraries, domain ontologies, distantly supervised relation extraction, or human-in-the-loop review).
- Keep raw text only on Chunk nodes. Documents carry metadata and provenance.
Step 4: Build a Vector Index
You can store embeddings in Neo4j or use a dedicated vector database. A simple, fast baseline is FAISS.
import faiss
import numpy as np
from neo4j import GraphDatabase
# Build an index from Chunk embeddings stored in Neo4j
vec_dim = 384 # all-MiniLM-L6-v2
index = faiss.IndexFlatIP(vec_dim)
chunk_ids = []
with driver.session() as sess:
result = sess.run("MATCH (c:Chunk) RETURN c.id as id, c.embedding as emb")
vectors = []
for rec in result:
chunk_ids.append(rec["id"])
v = np.array(rec["emb"], dtype=np.float32)
v /= np.linalg.norm(v) + 1e-12
vectors.append(v)
mat = np.vstack(vectors)
index.add(mat)
# Map from FAISS row -> chunk id
id_map = np.array(chunk_ids)
Step 5: Graph-Aware Retrieval
Combine semantic recall with topology-aware expansion. Here’s a practical pattern:
- Semantic seeds: top-k chunks by embedding similarity to the question
- Graph expansion: pull neighbors around those seeds—entities, cited documents, related topics, co-mentioned chunks
- Candidate assembly: gather chunks reached through the graph that likely carry corroborating evidence
- Re-ranking: cross-encoder or LLM scoring to select final N chunks
from sentence_transformers import CrossEncoder
cross = CrossEncoder("cross-encoder/ms-marco-MiniLM-L-6-v2")
def semantic_seeds(question, k=12):
q = embedder.encode([question])[0].astype(np.float32)
q /= np.linalg.norm(q) + 1e-12
D, I = index.search(q.reshape(1, -1), k)
return id_map[I[0]].tolist()
def graph_expand(seed_chunk_ids, max_candidates=200):
cypher = """
UNWIND $ids AS cid
MATCH (c:Chunk {id:cid})-[:MENTIONS|:ABOUT]->(e)
OPTIONAL MATCH (c)-[:CONTAINS]-(d:Document)
OPTIONAL MATCH (c)-[:ABOUT]->(t:Topic)
// one-hop neighborhood to related chunks via shared entity/topic/citations
OPTIONAL MATCH (c)<-[:CONTAINS]-(d)-[:CITES]->(d2:Document)-[:CONTAINS]->(c2:Chunk)
OPTIONAL MATCH (c)-[:MENTIONS|:ABOUT]->(e)<-[:MENTIONS|:ABOUT]-(c3:Chunk)
WITH DISTINCT c, collect(DISTINCT c2) + collect(DISTINCT c3) AS nbrs
UNWIND nbrs AS cand
WITH DISTINCT cand LIMIT $limit
RETURN cand.id AS id, cand.text AS text
"""
with driver.session() as sess:
recs = sess.run(cypher, ids=seed_chunk_ids, limit=max_candidates)
return [{"id": r["id"], "text": r["text"]} for r in recs]
def rerank(question, candidates, top_n=12):
pairs = [(question, c["text"]) for c in candidates]
scores = cross.predict(pairs)
ranked = sorted(zip(candidates, scores), key=lambda x: x[1], reverse=True)
return [c for c, _ in ranked[:top_n]]
def hybrid_retrieve(question, k_seeds=12, max_cands=200, top_n=12):
seeds = semantic_seeds(question, k=k_seeds)
cands = graph_expand(seeds, max_candidates=max_cands)
# include the seeds themselves to avoid missing high-similarity text
with driver.session() as sess:
seed_text = sess.run("MATCH (c:Chunk) WHERE c.id IN $ids RETURN c.id as id, c.text as text", ids=seeds)
for r in seed_text:
cands.append({"id": r["id"], "text": r["text"]})
# dedupe by id
uniq = {c["id"]: c for c in cands}.values()
return rerank(question, list(uniq), top_n=top_n)
This function returns a compact set of highly relevant chunks tied together by graph structure. These chunks are ready to be fed as grounded context to your LLM.
Step 6: Structured Queries with Cypher Templates
Many enterprise questions are inherently structured. For repeatable intents, define Cypher templates rather than always relying on natural-language-to-Cypher.
Examples:
- “Which authors collaborated with [person]?”
MATCH (p:Person {name:$name})<-[:MENTIONS]-(c1:Chunk)-[:CONTAINS]-(d:Document)-[:CONTAINS]->(c2:Chunk)-[:MENTIONS]->(co:Person)
WHERE co <> p
RETURN co.name AS collaborator, count(DISTINCT d) AS docs
ORDER BY docs DESC LIMIT 20;
- “Show documents that cite [document_id] and their topics”
MATCH (d:Document {id:$doc_id})<-[:CITES]-(citing:Document)
OPTIONAL MATCH (citing)-[:CONTAINS]->(:Chunk)-[:ABOUT]->(t:Topic)
RETURN citing.id AS id, citing.title AS title, collect(DISTINCT t.name) AS topics
LIMIT 50;
- “Top organizations co-mentioned with [topic]”
MATCH (:Topic {name:$topic})<-[:ABOUT]-(c:Chunk)-[:MENTIONS]->(o:Organization)
RETURN o.name AS org, count(*) AS freq
ORDER BY freq DESC LIMIT 20;
You can detect such intents with lightweight patterns or a classifier and route to the right template before falling back to hybrid retrieval.
Step 7: Assemble Grounded Prompts for the LLM
Concise, well-cited context prevents hallucinations. Include chunk IDs and document metadata.
def make_prompt(question, evidences):
header = "You are a careful analyst. Answer using only the EVIDENCE. Cite chunk ids like [c:ID]. If unsure, say you don't know.\n\n"
ev_txt = []
for e in evidences:
ev_txt.append(f"[c:{e['id']}] " + e['text'].strip())
context = "\n\n".join(ev_txt)
return f"{header}QUESTION: {question}\n\nEVIDENCE:\n{context}\n\nANSWER:"
You can use any chat completion API. Always track which chunks are used in the final answer for auditability.
Step 8: Evaluation—Don’t Skip This
Measure the pipeline with a small but representative test set.
Metrics:
- Faithfulness: percentage of answers fully supported by provided chunks (manual review or automated heuristics)
- Coverage/Recall: fraction of gold evidence chunks present in the retrieved set
- Precision: fraction of retrieved chunks that reviewers deem relevant
- Latency: end-to-end p95
- Context budget: average tokens used per query
Procedure:
- Create ~100–300 question–answer–evidence triples from your domain.
- Run baseline vector-only RAG.
- Run hybrid graph + vector retrieval.
- Compare faithfulness, coverage, and latency; iterate on schema, expansion rules, and re-ranking.
Operational Tips and Pitfalls
- Canonicalization: Merge duplicate entities. Use lowercase, trimmed names and alias lists. Consider string similarity thresholds (e.g., Jaro–Winkler) under human review.
- Provenance-first: Every edge should be explainable. Keep source doc IDs and stable references.
- Depth control: Limit expansion hops to 1–2 to cap latency and noise. Rely on re-ranking rather than deep traversals.
- Balance recall vs. precision: Start with generous recall (k≈10–20 seeds, 100–300 candidates) then tighten after measuring precision.
- Incremental updates: Upsert by document ID. Maintain a change log for re-embedding only changed chunks.
- Safety and privacy: Filter PII; restrict sensitive nodes/edges by role. Apply row-level security where possible.
- Monitoring: Track query types, cache hit rates, token usage, and any “empty context” failures.
Extending the Tutorial
- Add richer relations: WORKS_AT, AUTHORED, PUBLISHED_IN, LOCATED_IN, DERIVES_FROM.
- Use a graph algorithm: Personalized PageRank or community detection to identify influential nodes around a query topic.
- NL-to-Cypher: Train a few-shot prompt or a constrained decoder that emits only allowed labels and relations from your schema.
- Multi-index retrieval: Blend BM25 keyword search, vector search, and graph paths; then learn weights via logistic regression on your eval set.
- Reranking upgrades: Try larger cross-encoders or LLM-based re-rankers in a two-stage cascade.
End-to-End Example
Putting it together for a single question:
question = "Which organizations frequently co-occur with the topic 'Generative AI' and what sources cite them?"
seeds = semantic_seeds(question, k=15)
candidates = graph_expand(seeds, max_candidates=250)
# Add seeds and dedupe
with driver.session() as sess:
seed_text = sess.run("MATCH (c:Chunk) WHERE c.id IN $ids RETURN c.id as id, c.text as text", ids=seeds)
for r in seed_text:
candidates.append({"id": r["id"], "text": r["text"]})
uniq = {c["id"]: c for c in candidates}.values()
final_ctx = rerank(question, list(uniq), top_n=15)
prompt = make_prompt(question, final_ctx)
print(prompt[:1000]) # send to your LLM of choice
Optionally, add a structured pass first:
// Top orgs tied to a topic with supporting docs
MATCH (:Topic {name:$topic})<-[:ABOUT]-(c:Chunk)-[:MENTIONS]->(o:Organization)
WITH o, count(*) AS freq
ORDER BY freq DESC LIMIT 10
MATCH (o)<-[:MENTIONS]-(c2:Chunk)<-[:CONTAINS]-(d:Document)
RETURN o.name AS org, freq, collect(DISTINCT d.id)[..5] AS evidence_docs;
What You’ll Have by the End
- A working KG-backed retriever that preserves structure and provenance
- A reproducible ingestion pipeline and vector index
- Measurable improvements in faithfulness and answerability on structured questions
Next Steps
- Enrich schema and relations driven by your evaluation findings.
- Introduce caching for frequent queries (semantic + graph sub-results).
- Productionize: containerize the retriever, move to managed graph/vector services, add tracing and dashboards.
- Expand your evaluation set and run regular regressions before each schema or model change.
With these building blocks, you’ve moved from “semantic search plus LLM” to a principled, graph-aware retrieval system that can explain its answers and scale with your domain’s complexity.
Related Posts
A Practical Guide to Multi‑Modal RAG: Images Plus Text, End‑to‑End Tutorial
Build a practical multi‑modal RAG system that retrieves from images and text using OCR, captions, CLIP embeddings, and vector search.

Integrating an AI Writing Assistant via API: Architecture, Code, and Best Practices
A practical guide to integrating an AI writing assistant via API—architecture, prompt design, code samples, safety, evaluation, and performance optimization.

Open-Source LLM Deployment Guide: From Laptop Prototype to Production
Practical, end-to-end guide to deploying open-source LLMs—from model choice and hardware sizing to serving, RAG, safety, and production ops.
